-
PDF
- Split View
-
Views
-
Cite
Cite
Chloé Marchive, Rim Mzid, Laurent Deluc, François Barrieu, Julien Pirrello, Adrien Gauthier, Marie-France Corio-Costet, Farid Regad, Bernard Cailleteau, Saïd Hamdi, Virginie Lauvergeat, Isolation and characterization of a Vitis vinifera transcription factor, VvWRKY1, and its effect on responses to fungal pathogens in transgenic tobacco plants, Journal of Experimental Botany, Volume 58, Issue 8, June 2007, Pages 1999–2010, https://doi.org/10.1093/jxb/erm062
Close - Share Icon Share
Abstract
Pathogen attack represents a major problem for viticulture and for agriculture in general. At present, the use of phytochemicals is more and more restrictive, and therefore it is becoming essential to control disease by having a thorough knowledge of resistance mechanisms. The present work focused on the trans-regulatory proteins potentially involved in the control of the plant defence response, the WRKY proteins. A full-length cDNA, designated VvWRKY1, was isolated from a grape berry library (Vitis vinifera L. cv. Cabernet Sauvignon). It encodes a polypeptide of 151 amino acids whose structure is characteristic of group IIc WRKY proteins. VvWRKY1 gene expression in grape is regulated in a developmental manner in berries and leaves and by various signal molecules involved in defence such as salicylic acid, ethylene, and hydrogen peroxide. Biochemical analysis indicates that VvWRKY1 specifically interacts with the W-box in various nucleotidic contexts. Functional analysis of VvWRKY1 was performed by overexpression in tobacco, and transgenic plants exhibited reduced susceptibility to various fungi but not to viruses. These results are consistent with a possible role for VvWRKY1 in grapevine defence against fungal pathogens.
Introduction
Plants are exposed to various pathogens in nature. To counter microbial attack, they have evolved multiple and complex defence strategies. Most of these microbes can be dealt with by the plant basal defence system that limits the growth of non-host pathogens. Plant disease resistance can be induced via host recognition of pathogen elicitors (also designated as pathogen-associated molecular patterns or PAMPs; Nürnberger et al., 2004). Bacterial flagellin is an excellent example of a general elicitor that induces resistance through interaction with a plasma membrane-localized receptor kinase (Zipfel et al., 2004). This mode of pathogen recognition resembles pathways of innate immunity response (Gomez-Gomez and Boller, 2002; Navarro et al., 2004). The plant innate immune response is initiated when a plant resistance (R) gene product either directly or indirectly recognizes specific effector molecules or proteins produced by pathogen avirulence (Avr) genes (Martin et al., 2003). This type of recognition triggers defined signal transduction cascades within the infected plant cell, leading to the generation of endogenous signalling coumpounds and to the subsequent biosynthesis of antimicrobial proteins locally as well as systemically, i.e. in distant parts of the plant, a phenomenon termed systemic acquired resistance (SAR) (Ryals et al., 1996). Often, plant resistance is accompanied by rapid localized and programmed cell death at the infection site, termed the hypersensitive response (HR), which is preceded by physiological disturbances (ion fluxes, pH changes, membrane depolarization, and oxidative burst; Nürnberger and Scheel, 2001). HR and SAR are accompanied by the accumulation of defence molecules and proteins, resulting in cell wall modifications and the production of antimicrobial molecules (phytoalexins) and a large group of pathogenesis-related (PR) proteins, some of which have also antimicrobial activities (Van Loon and Van Strien, 1999; Narasimhan et al., 2005).
Elicitor- or pathogen-activated transcription factors play an important role in controlling defence gene expression and plant resistance responses. Five major families of plant transcription factors (bZIP, WRKY, MYB, EREBF, and homeodomain proteins) have been shown to participate in the regulation of plant defence responses (Rushton and Somssich, 1998). It is generally assumed that WRKY transcription factors act as major regulatory proteins by binding to the W-box, a common promoter element contained in several SAR gene promoters (Maleck et al., 2000). This class of transcription factors belongs to a large family of proteins mainly present in plants, and is characterized by their highly conserved DNA-binding region termed the WRKY domain. In plants, many WRKY proteins are involved in the defence against attack by phytopathogens such as bacteria (Dellagi et al., 2000; Asai et al., 2002; Chen et al., 2002; Deslandes et al., 2002; Dong et al., 2003), fungi (Beyer et al., 2001; Asai et al., 2002; Chen et al., 2002; Kalde et al., 2003), and viruses (Wang et al., 1998; Yang et al., 1999; Chen et al., 2002; Yoda et al., 2002; Liu et al., 2004). Furthermore, a role in various physiological processes has also been suggested, including embryogenesis, seed coat and trichome development, senescence, regulation of biosynthetic pathways, and hormonal signalling (Ulker and Somssich, 2004; Lagace and Matton, 2004; Xu et al., 2004; Zhang et al., 2004; Zou et al., 2004; Xie et al., 2005).
Cultivated grapevines (Vitis vinifera L.) are susceptible to many pathogens such as phytoplasmas, viruses, bacteria, and fungi (Galet, 1996; Martini et al., 2002; Ferreira et al., 2004). Among them, the most important are grey mould (Botrytis cinerea), powdery mildew (Erysiphe necator), and downy mildew (Plasmopara viticola), which cause extensive loss of quantity and quality in harvested berries. Consequently, wine growing requires extensive use of phytochemicals. For instance, in France, 50% of the total mass of these products are used in vineyards, which represent only 3.7% of farmed surfaces. Intensive use of chemicals has led to the development of microorganisms resistant to certain types of fungicides, and some products are now prohibited due to their high toxicity. Nowadays, several research strategies are being pursued so that wine growers may produce healthy fruits right up to maturity with a minimum use of chemical treatments.
The aim of this study was to identify components of the grape defence mechanisms, and we focus on a WRKY transcription factor expressed in grape berries because fruits, rich in sugar and other nutrients, provide an ideal target for pathogens. To this end, a full-length cDNA, VvWRKY1, was isolated from a Vitis vinifera grape berry library. The ability of the corresponding protein to bind specifically to W-box DNA elements was demonstrated. Then, its expression was characterized during the development of healthy plants, berries, and leaves, and under stress conditions. The biological role of VvWRKY1 was assessed by its overexpression in Nicotiana tabacum. These transgenic plants exhibited decreased susceptibility towards several fungal pathogens compared with the wild type, implicating this transcription factor in plant defence responses.
Material and methods
Grape plant material
During the 2000 growing season, grape berries from V. vinifera L. cv. Cabernet Sauvignon plants were collected from the ‘Domaine du Grand Parc’ (INRA, Latresne, France), at different stages of maturity [from 20 to 107 days after flowering (daf)]. Separate seeds and skin/flesh samples were immediately frozen in liquid nitrogen before storage at −80 °C.
Rooted plants obtained from grape cuttings (V. vinifera L. cv. Cabernet Sauvignon) were cultivated on a sandy soil in a growth room programmed for 25/20 °C under a 16/8 h light/dark photocycle and at 75% relative humidity (Ollat, 1998). Leaves were collected at different stages, from apices and very young leaves (2–3 cm wide) to older leaves (9–10 cm wide), to study VvWRKY1 expression in these organs.
Tobacco plants (N. tabacum cv. Xanthi) were grown in vitro at 25 °C on MS medium (Murashige and Skoog, 1962) supplemented with kanamycin (100 mg l−1) for transgenic lines, or without kanamycin for control plants, under a 16 h photocycle at 25/20 °C. Plantlets were transferred to a greenhouse to produce seeds.
VvWRKY1 cDNA isolation, vector construction, and plant transformation
The VvWRKY1 full-length cDNA was cloned from a grape berry cDNA library (V. vinifera L. cv. Cabernet Sauvignon veraison stage; Deluc et al., 2006) by PCR screening with oligonucleotides designed in the consensus WRKY sequence (F, 5′-TGGMGDAARTAYGGRCGAAR-3′; and R, 5′-YTTCGYCCRTAYTTHCKCCA-3′). These primers were used in combination with T7 and 5′ primers located in the pTriplex vector (Smart cDNA library construction kit, Clontech, Palo Alto, CA, USA). After cloning in pGEM-T Easy plasmid (Promega, Charbonnières, France), PCR products were sequenced (Genome Express, Meylan, France), and specific oligonucleotides defined within the 5′ and 3′ non-coding regions were used to amplify VvWRKY1 full-length cDNA.
To overexpress VvWRKY1 in tobacco, XbaI and SacI restriction sites were added, respectively, at the 5′ and 3′ ends of the cDNA by PCR amplification with modified primers (F, 5′-CTCAATCTAGATCAGTCTCTC-3′; and R, 5′-TCCAAGAGCTCATATGGGTGT-3′). The full-length modified fragment was inserted into a pGEM-T Easy vector and, after restriction with XbaI and SacI, subcloned into the same restriction sites of the binary p35SGiBin19 plasmid (Karimi et al., 2000). The resulting p35SGiBin19-VvWRKY1 containing the full coding sequence of VvWRKY1 in the sense orientation downstream of the 35S promoter was first introduced into Agrobacterium tumefaciens LBA 4404 strain (Hoekema et al., 1983). Tobacco (N. tabacum cv. Xanthi) was then transformed using the leaf disc method (Horsch et al., 1985). Transgenic progeny lines were selected on MS medium supplied with kanamycin (100 mg l−1) and carried to homozygosity.
In vitro VvWRKY1 protein synthesis and electrophoretic mobility shift assay
The VvWRKY1 protein was produced by the in vitro transcription and translation method with the TnT T7 quick system for the PCR DNA system (Promega, Charbonnières, France) according to the manufacturer's instructions. The coding sequence was first amplified with Turbo-Pfu (Stratagene) using the following primer pairs: F, 5′- AGATCCTAATACGACTCACTATAGGGAGCCACCATGAAGGCCACCAAATACTT-3′; and R, 5′- (T)32GGTGAAATGGAAACATTCAT-3′, and the PCR product was used as template. Protein detection was performed using the Transcend™ non-radioactive translation detection system (Promega, Charbonnières, France). Double-stranded probes were labelled with [33P]ATP using a 5′ end labelling kit (Amersham Pharmacia Biotech, Buckinghamshire, UK) and purified on a 6% non-denaturing polyacrylamide gel in 0.5× TBE buffer at 250 V. The resultant 120 000 cpm labelled DNA probe was incubated with 4 ml of protein reaction mixture, 2 μg of poly(dI–dC)–poly(dI–dC) and the binding buffer (20 mM TRIS, 50 mM NaCl, 10% glycerol, 7 mM β-mercaptoethanol) in a 20 μl reaction volume. In competition experiments, specific competitor was added in 200-fold molar excess prior to the addition of the protein. The mixture was incubated for 30 min at room temperature and analysed on a non-denaturing 6% polyacrylamide gel in 0.5× TBE buffer at 150 V. The gel was then dried and autoradiographed using a Fujix BAS 2000 system (Fuji Photofilm, Tokyo, Japan).
Wounding and chemical treatments
Two-month-old rooting plants obtained from grape cuttings cultivated in a growth room were used for treatments. For wounding experiments, four fully developed leaves were cut on four sites with scissors and pooled for each time point. For chemical treatments, plants were sprayed with 5 mM salicylic acid (SA), 10 mM ethephon (2-chloroethylphosphonic acid), or 10 mM H2O2. Mock treatments were performed by spraying plants with water. Leaves from three plants were pooled for each time point, frozen in liquid nitrogen, and stored at −80 °C before RNA extraction. All experiments were conducted at least twice. The chemicals were purchased from Sigma (St Louis, MO, USA).
RNA isolation and RT-PCR analysis
Total RNA was isolated from grapevine by the LiCl precipitation method (Asif et al., 2000) and from N. tabacum using the TriReagent method (Molecular Research Center, Cincinnati, OH, USA), according to the manufacturer's instructions. The first-strand cDNA was synthesized from 2 μg of DNase-treated RNA using M-MLV reverse transcriptase (Promega, Madison, WI, USA). To ensure that the PCR products were generated from cDNA and not genomic DNA, proper control reactions were carried out without reverse transcriptase treatment.
PCR amplifications on grape templates were performed using primers specific for the VvWRKY1 gene (F, 5′-GAGAATGATATGGAAAGAGTGG-3′; and R, 5′-CATTCGTTCTCAGACACAATA-3′), the β-1,3 glucanase gene (F, 5′-GCACTCGCATCAGCAGGC-3′; and R, 5′-GTCACTCTTGAGGGATTC-3′; accession no. AF239617), and the elongation factor EF1γ gene (F, 5′-GCGGGCAAGAGATACCTCAA-3′; and R, 5′-TCAATCTGTCTAGGAAAGGAAG-3′; accession no. AF176496). The number of PCR cycles was 30, 25, and 27 cycles for VvWRKY1, β-Glu, and EF1γ, respectively. The tobacco transgene was amplified using specific primers (F, 5′-CCTTTTCCACCAAACATG-3′; and R, 5′-TCATCGCAAGACCGGCAACA-3′) with 30 PCR cycles. The Ubiquitin mRNA was amplified as tobacco constitutive control (F, 5′-TCCAGGACAAGGAGGGTAT-3′; and R, 5′-GAGACCTCAGTAGACAAAGC-3′; accession no. U66264) with 23 cycles. For each gene, the PCR cycle number chosen was in the log-linear phase of the amplification cycle. RT-PCR products were separated on 1.6% agarose gels, BET stained, and quantified by densitometry using Quantity One software (Bio-Rad). To confirm RT-PCR specificity, the products were inserted into pGEM-T Easy vector (Promega, Charbonnières, France) and subsequently sequenced.
Pathogen challenge of transgenic tobacco
For Pythium F (strain 00PR201, from INRA Bordeaux, France) susceptibility studies, mycelium gelose discs from a 3-d-old culture at 20 °C on V8 medium [V8 vegetable juice 20% (v/v), CaCO3 0.25% (w/v), and agar 17 g l−1) were placed in the crown area of 20 seedlings (3 weeks old) per line. The assay was carried out in Petri dishes containing Whatman paper saturated with water. For each line, a control corresponding to the same number of plantlets inoculated with a disc of V8 agar medium without mycelium was performed. Infected plants were grown in a growth room at 20 °C under a 16 h photocycle. Symptoms were estimated at 7, 10, and 15 d post-inoculation (dpi) according to a scale from 0 to 7 depending on root necrosis development: 0=no symptoms, 1–9=percentage of the root surface that is necrotic; 1=less than 20%, 2=20%, 3=between 30 and 40%, 4=60%, 5=between 60 and 80%, 7=more than 80%, and 9=100%.
For disease susceptibility studies with Peronospora tabacina (downy mildew), Erysiphe cichoracearum (powdery mildew), and potato virus Y (PVYN), strains and inoculum preparations were provided by the Institut du Tabac de Bergerac (ITB, France). For each assay, symptoms were observed on cultivars with different levels of susceptibility (provided by ITB) to validate the progress of the disease. T2 progeny of transgenic lines were analysed and the untransformed N. tabacum cv. Xanthi line was used as control. Seeds were germinated on a blotting paper and transferred after 7 d (two cotyledons-stage) to the greenhouse before inoculation.
For downy mildew (P. tabacina) infections, six plants (10 weeks old) per line were transferred to the growth room (10/14 h light/dark cycle at 22/16 °C). Two leaves per plant were infiltrated on two sites with 100 μl of a sporangia suspension (100 000 spores ml−1) using a needleless syringe. Disease intensity was evaluated 9 dpi by measuring the development of chlorotic flecks around the inoculation site. A second scale was used to estimate the fungal sporulation: 0=no visible symptoms, 1–3=conidia barely detectable, 4–6=moderate amount of conidia production, 7–9=high amount of conidia.
To analyse susceptibility to the powdery mildew agent (E. cichoracearum), 200–500 conidia (as controlled under the microscope) were applied with a cotton tip to two sites of the upper side of two leaves per plant. Twelve plants per line (10 weeks old) were inoculated and maintained in a growth chamber at 23/16 °C under a 10 h photoperiod. After 10 d and 13 d, the percentage of leaf surface covered by powdery mildew mycelium was estimated.
Tobacco plants (4 weeks old) were inoculated with PVYN by rubbing fully expanded leaves with wet carborundum plus PVYN in a 50 mM phosphate buffer, pH 7.0. Two leaves per plant and six plants per line were infected. Plants were maintained in a greenhouse at 20/25 °C. Symptoms were observed 21 d after infection and noted on a scale from 1 to 11 depending on necrotic and mosaic leaf symptom development: 0=no visible symptoms, 3=slight vein clearing on infected leaves, 5=extent of the veins clearing on a few leaves, 7=veinlets and veins become necrotic, 9=vein necrosis extends to at least five leaves, 11=all leaves develop vein necrosis symptoms.
All the data were statistically analysed by analysis of variance (ANOVA) test (P-value <0.05 or <0.01).
Results
Cloning of a grapevine cDNA encoding a group IIc WRKY transcription factor
Degenerate oligonucleotide primers corresponding to the conserved core sequence of WRKY proteins (WRKYGQK) were used to screen a cDNA library prepared from grape berries harvested at veraison stage (V. vinifera L. cv. Cabernet Sauvignon). One of the overlapping sequences obtained after cloning and sequencing of the amplified products corresponded to a full-length cDNA of 849 bp and was designated VvWRKY1. It contains a 68 bp 5′-untranslated region (UTR) and a 298 bp 3′ UTR with a poly(A) tail (Fig. 1). The predicted open reading frame encodes a protein of 151 residues with a predicted molecular mass of 17.7 kDa. Moreover, analysis of the VvWRKY1 protein sequence using the PSORT program (Nakai and Kanehisa, 1992, http://psort.nibb.ac.jp) revealed the presence of a putative nuclear localization signal, RKHR, between positions 56 and 59 of the amino acid sequence (Fig. 1).
cDNA and deduced amino acid sequence of VvWRKY1. The 5′ and 3′ UTRs are in italics. The WRKY motif is shown in bold, the two cysteines and the two histidines of the zinc-finger motif are circled, and the putative nuclear localization signal (RKHR) is labelled with a rectangle. The first number written on the top of the sequence corresponds to the nucleotide number from the initiator ATG and the second one to the amino acid number. The sequence has been deposited in the GenBank database under accession no. AY585679.
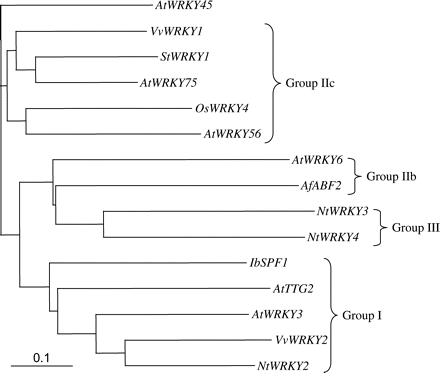
WRKY proteins are divided into three main classes based on sequence similarity. Database searches showed that the predicted VvWRKY1 polypeptide is highly similar to group II WRKY proteins, containing only one WRKY domain at its N-terminal end followed by a Cys2/His2-type zinc-finger motif. Additional structural motifs allowed the distinction of different subgroups (Eulgem et al., 2000), and VvWRKY1 shows a high degree of similarity to the subgroup IIc (Fig. 2). VvWRKY1 is closely related to Arabidopsis thaliana AtWRKY75 (Eulgem et al., 2000) and Solanum tuberosum StWRKY1 (Dellagi et al., 2000) which share 59% and 53.9% amino acid sequence identities, respectively.
VvWRKY1 belongs to group II of the plant WRKY family. Phylogenetic tree of a selection of WRKY proteins. The deduced amino acid sequence of VvWRKY1 was aligned with those encoded by 14 genes in GenBank using the CLUSTALX program (Thomson et al., 1997). The phylogenetic tree was created using the TreeView program (Page, 1996). Accession nos: AfABF2, CAA88331; AtWRKY3, AAK28311; AtWRKY6, AAK28312; AtWRKY45, AAL29428; AtTTG2 (WRKY44), AAM61951; AtWRKY56, AAL61858; AtWRKY75, AAL50784; IbSPF1, BAA06278; NtWRKY2, AAD16139; NtWRKY3, AAF61863; NtWRKY4, BAA86031; OsWRKY4, DAA05069; StWRKY1, AAU50687; VvWRKY2, AAT46067. The scale bar at the bottom left displays a distance corresponding to 0.1 amino acid substitutions per site.
Otherwise, comparison of the VvWRKY1 sequence with expressed sequence tags (ESTs) published in the Grape TIGR database indicates that it corresponds to the TC50592 sequence identified from an abiotic stressed grapeberry library (V. vinifera L. cv. Chardonnay). The other two related TC sequences also come from abiotic stressed berry or leaf libraries (TC41321 and TC46341) and exhibit a very high sequence identity with VvWRKY1. The TIGR database does not contain any ESTs from pathogen-infected grapevine libraries.
VvWRKY1 binds to W-box elements
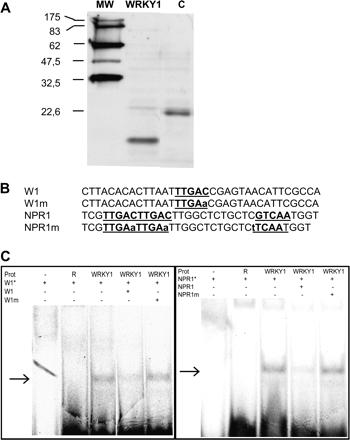
WRKY proteins have been identified as transcription factors that bind DNA sequences containing W-boxes (TGAC). In order to test VvWRKY1 protein–W-box DNA interactions, electrophoretic mobility shift assays (EMSAs) were performed. Production of recombinant VvWRKY1 in bacterial (Escherichia coli) and yeast (Saccharomyces cerevisiae) systems proved unsuccessful, most probably due to toxicity effects (data not shown). Therefore, the protein was synthesized by an in vitro transcription and translation assay. The biotinylated protein band of VvWRKY1 was detected by a chemiluminescent method (Fig. 3A).
Specific W-box binding activity of VvWRKY1. (A) Detection of the in vitro synthesized VvWRKY1 protein. A 2 μl aliquot of the reagent was used for SDS-PAGE. Separated proteins were transferred onto a nitrocellulose membrane and detected using a chemiluminescent method. MW corresponds to the molecular ladder, WRKY1 to the VvWRKY1 protein, and C is an in vitro synthesized control protein of 22.6 kDa. (B). Nucleotide sequences and their mutant versions used as probes and specific competitors in EMSA. W1 was derived from the PcPR1 promoter (Rushton et al., 1996) and NPR1 is from the NtNPR1 promoter (Yu et al., 2001). The W-box elements (TGAC core) are underlined. The mutated nucleotides are in lower case. (C). Binding activity of VvWRKY1 to the W-box element was determined by gel mobility shift assay. Labelled probes (W1*, NPR1*) were incubated with VvWRKY1 protein (WRKY1). Competitors (W1, NPR1, or mutated probes) were added in 200-fold molar excess. The second lane represents DNA binding incubation with the rabbit reticulocyte lysate (R). Arrows indicate DNA–protein complexes.
In the absence of known grape target genes, EMSA was performed with three oligonucleotide probes previously shown to be bound by WRKY proteins in other plant species. These sequences contain W-boxes in different arrangements and with different surrounding sequences. Two sequences, W1 and NPR1, were used. The first one, W1, derived from the parsley PR1 promoter contains a single W-box TTGAC (Fig. 3B; Rushton et al., 1996). The second sequence is derived from the A. thaliana NPR1 gene promoter (Yu et al., 2001) and is composed of three W-box sequences (TTGAC) within 28 bp (two W-boxes in tandem and a third W-box in the reverse orientation; Fig. 3B).
A shift band was observed with each probe incubated with the in vitro transcribed and translated protein (Fig. 3C). To verify VvWRKY1 binding specificity, competition experiments were conducted by adding unlabelled competitors in 200-fold molar excess to the binding assays. Competition with the unlabelled specific probe strongly reduced the binding signal, whereas addition of an unlabelled W-box-mutated probe (changing TTGAC to TTGAA) to the reaction volume had no effect. Similar binding specificities on the TTGAC sequence were seen with the promoter probes W1 and NPR1. These results demonstrate that the TTGAC core sequence is necessary for binding of VvWRKY1 protein to these DNA probes.
VvWRKY1 expression is developmentally regulated
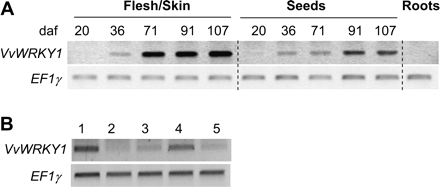
Total RNA was isolated from various V. vinifera L. cv. Cabernet Sauvignon tissues and the expression of VvWRKY1 was investigated by semi-quantitative RT-PCR using primers designed within the 3′ UTR (Fig. 4). VvWRKY1 was not expressed in roots but is expressed in fruit and leaves. In both of these latter organs, RT-PCR analysis revealed that expression of this transcription factor gene was developmentally controlled. The VvWRKY1 RNA level was very low in berries before veraison (about 60 daf), but it increased significantly after this stage and kept on accumulating throughout ripening (Fig. 4A). This increase appeared to be greater in flesh/skin than in seeds. In leaves, accumulation of VvWRKY1 transcripts was high in very young leaves and apices and in well-developed leaves (Fig. 4B). In contrast, in young leaves and in mature leaves, a lower signal was observed.
VvWRKY1 expression during grape berry and leaf development. (A) Berries were harvested 20, 36, 71, 91, and 107 days after flowering (daf). Flesh and skin were separated from seeds. Semi-quantitative RT-PCR analyses were performed on total RNA prepared from these samples for VvWRKY1 expression. Elongation factor EF1γ was used as quantitative control. (B) Leaves of grape cuttings were harvested at different stages of leaf development. Numbers above the lines correspond to: 1, apex and very young leaves; 2, young leaves of about 2–3 cm wide; 3, 4–5 cm wide leaves; 4, 6–7 cm wide leaves; 5, mature and well-developed leaves about 9–10 cm wide.
These results indicate that expression of VvWRKY1 is not fruit specific and is strongly regulated during development in fruits and leaves.
VvWRKY1 expression is regulated by defence signals
The effect of wounding and some of the signal molecules known to trigger plant defence gene expression were investigated on the expression of VvWRKY1. Leaves of grapevine cuttings were wounded using scissors or sprayed with solutions containing SA, ethephon, or H2O2, and were collected for RNA isolation at the time points indicated. VvWRKY1 expression was analysed by semi-quantitative RT-PCR. The expression of β-Glu encoding a pathogenesis-related protein (PR2) was monitored as a plant defence control. Two independent experiments were performed and gave similar results.
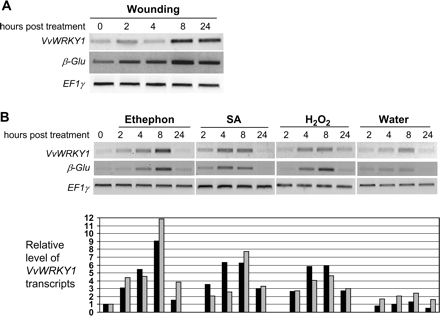
After wounding, the level of VvWRKY1 transcripts increased progressively from 2 h and reached a maximum at 8 h (Fig. 5A). This activation persisted 24 h post-treatment (hpt). The same pattern is observed for β-Glu gene expression.
VvWRKY1expression in response to wounding (A) and defence signal molecules (B). Leaves from grape cuttings were wounded by scissors or sprayed with 5 mM SA, 10 mM ethephon, or 10 mM H2O2. Control plants were sprayed with water. Leaves were harvested at the indicated times after treatment for preparation of total RNA. Accumulation of VvWRKY1 and β-Glu (a β-1,3 glucanase gene) transcripts was monitored by semi-quantitative RT-PCR. EF1γ amplification was used as constitutive control. VvWRKY1 transcript accumulation was quantified by densitometry using Quantity One software (Bio-Rad), for two independent biological experiments (black columns, first assay; and grey columns, second assay); each value was corrected according to the EF1γ amplification signal and expressed as a ratio between the corrected value and the control time point (at 0 h) corrected value.
Furthermore, it was observed that all signal molecules used induced an increase of VvWRKY1 and β-Glu expression from 2 hpt. The expression of VvWRKY1 and β-Glu was weakly affected by water (Fig. 5B), indicating that spraying itself did not affect gene expression. On the other hand, the strong transient induction of β-Glu expression observed 8 hpt for all compounds used confirms the efficiency of the treatments. The strongest induction of VvWRKY1 expression was detected 8 h after spraying the plants with ethephon, an ethylene-releasing compound. After SA or H2O2 treatments, transcript accumulation commenced at 2 hpt and reached a maximum at 4 hpt and 8 hpt before decreasing at 24 hpt. In each case, the activation of VvWRKY1 expression was transient, with VvWRKY1 mRNA levels decreasing at 24 hpt. It is also interesting to note that VvWRKY1 and β-Glu expression patterns appeared quite similar in response to the treatments performed in this study.
These results show that VvWRKY1 expression is positively affected by several defence signalling compounds and by wounding.
Ectopic expression of VvWRKY1 in tobacco results in reduced fungal susceptibility
The putative role of VvWRKY1 in plant defence was also addressed by overexpressing the cDNA under control of the 35S promoter of cauliflower mosaic virus in N. tabacum cv. Xanthi plants, since grapevine transformation still presents several difficulties. None of the generated transgenic plants showed any phenotypical changes compared with control plants. The expression level of the transgene was estimated for 10 independent lines by semi-quantitative RT-PCR using specific primers. Three lines were chosen on the basis of different accumulation levels of VvWRKY1 transcripts, two lines sharing a high level (lines 10 and 18) and one line (17) having a low level of transgene expression (Fig. 6A). Southern blot analysis revealed one copy of the transgene for line 10 and at least three copies for lines 17 and 18 (data not shown).
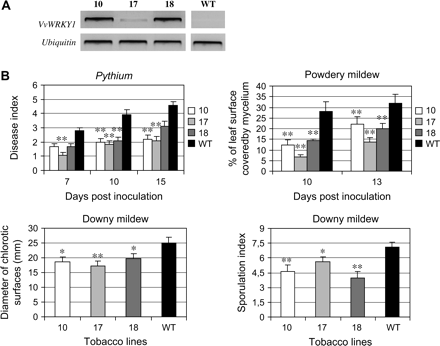
Disease tolerance analysis of VvWRKY1-overexpressing tobacco lines. (A) Expression levels of the transgene in the three VvWRKY1-overexpressing tobacco lines were estimated by RT-PCR using total RNA prepared from four leaves from three in vitro grown T2 homozygous plants for each line. Ubiquitin amplification was used as constitutive control. WT indicates the wild-type tobacco line. (B) Transgenic tobacco susceptibility to different fungal pathogens. The three selected transgenic tobacco lines (10, 17, and 18) and wild-type plants (WT) were inoculated with Pythium, downy mildew, or powdery mildew. Disease index (dependent on the root necrosis level) or leaf surface covered by mycelium were estimated for Pythium and powdery mildew, respectively, at different time points after inoculation. For downy mildew, two types of symptoms were evaluated. Fungal growth was estimated 6 d post-inoculation by measuring the diameter of the chlorotic surface for each inoculation site. The sporulation index (as described in the Materials and methods) of the fungus was also reported 9 d post-inoculation. Data represent means, and bars indicate standard errors. Data were statistically analysed by ANOVA, and asterisks indicate values that are significantly different (*P <0.05; **P <0.01) from the wild type.
Because many WRKY transcription factors have been shown to be involved in the activation of defence gene transcription, and particularly of PR genes (Yu et al., 2001), PR gene expression was investigated in the VvWRKY1-overexpressing plants under normal growth conditions. Despite the study of three generations of transgenic plants, no clear correlation between the transgene expression level and the amount of transcripts encoding PR proteins (five classes of PR proteins have been tested: PR1, PR2, PR3, PR4, and PR8) could be established. Molecular analysis by semi-quantitative RT-PCR showed that expression of some PR genes was slightly increased in only some lines compared with control plants (data not shown). These results suggest that VvWRKY1 ectopic expression does not act directly on the regulation of PR gene expression.
Susceptibility tests were performed on T2 progeny of these three transgenic lines using several types of fungi and one virus, PVYN. Pathogenic fungi with particular agronomic interest for tobacco were chosen: two oomycetes, Pythium (causing plantlets damping off) and P. tabacina (the tobacco downy mildew agent), and an ascomycete, E. cichoracearum (the tobacco powdery mildew agent). Susceptibility was estimated for each disease at different time points post-infection (Figs 6B, 7). Results were submitted for statistical analyses (ANOVA, P-value <0.05) to reveal their significance.
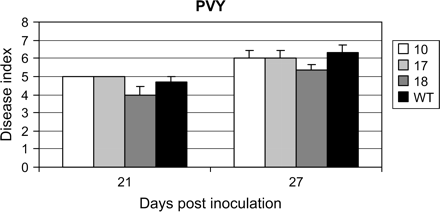
Susceptibility levels of VvWRKY1 transgenic tobacco lines infected by potato virus Y. Three transgenic tobacco lines (10, 17, and 18) and wild-type plants (WT) were inoculated with PVYN. Necrotic and mosaic symptoms on infected leaves were evaluated 21 d and 27 d after infection according to a visual scale from 1 to 11. Data represent means, and bars indicate standard errors. Data were statistically analysed by ANOVA, and asterisks indicate values that are significantly different (P <0.05) from the wild type.
For tolerance to Pythium, disease symptoms were estimated by visual scoring according to a severity scale at 7, 10, and 15 dpi. All transgenic lines showed reduced symptoms, with a 3-fold reduction of necrotic root surface in VvWRKY1 plants compared with wild-type plants. Indeed, at 10 dpi, transgenic lines developed necrosis on 20% of total root surfaces (disease index 2) in comparison with 60% for wild-type plants (disease index 4). The three transgenic lines were also more tolerant to powdery mildew, the percentage of leaf surface covered by mycelium 10 and 13 dpi being reduced from 1.5-fold to 4-fold (line 18 at 13 dpi and line 17 at 10 dpi, respectively) compared with the wild type (Fig. 6B). Concerning downy mildew infection experiments, two types of symptoms are shown in Fig. 6B: the diameter of chlorotic surfaces at 6 dpi and the sporulation rate at 9 dpi. For each observation, the three transgenic lines were slightly but significantly less affected by this pathogen than the control tobacco. Necrotic symptoms were also observed at infection sites at a significantly higher level than in control plants (data not shown). As a consequence of the limited fungal growth and the presence of necrotic sites, fungal sporulation 9 dpi was significantly reduced in the transgenic plants. Interestingly, it can be noted that a weak level of VvWRKY1 expression (line 17) was sufficient to induce a significantly better tolerance to theses fungal pathogens.
Disease resistance of the transformed tobacco plants against a virus, PVYN, was also studied. As shown in Fig. 7, no difference was observed between wild-type and transgenic plants.
Taken together, these data suggest the involvement of the VvWRKY1 transcription factor in activating defence mechanisms toward fungal pathogens in tobacco.
Discussion
Despite the existence of numerous links between WRKY transcription factors and plant defence mechanisms, direct evidence for the involvement of WRKY proteins in this process remains limited. In most cases, their expression is up-regulated in pathogen-infected or SA-treated plants. This up-regulation is often correlated with pathogen resistance as they are expressed specifically during incompatible reactions (Liu et al., 2004; Park et al., 2006). Functional studies have only been performed in A. thaliana for a few WRKY transcription factors. To date, two Arabidopsis WRKY factors (AtWRKY22 and AtWRKY29) have been identified as important downstream compounds of the MAPK pathway that confer resistance to both bacterial and fungal pathogens (Asai et al., 2002). Overexpression of AtWRKY70 increased resistance to virulent pathogens (Li et al., 2004, 2006). Recently, a WRKY transcription factor (OsWRKY71) was overexpressed in rice, and transgenic plants showed an enhanced resistance to virulent bacteria (Liu et al., 2006). Similar results have been shown with Arabidopsis plants expressing AtWRKY18, albeit in a development-dependent manner (Chen and Chen, 2002). However, nearly all such lines showed altered leaf morphologies and changes in flowering time (Chen and Chen, 2002; Robatzek and Somssich, 2002; Li et al., 2004).
In the present study, a cDNA, designated VvWRKY1, encoding a polypeptide of 151 amino acids, was identified. Phylogenetic analysis showed that VvWRKY1 belongs to the subgroup IIc (Dong et al., 2003), with StWRKY1 and AtWRKY75 (corresponding to AtWRKY64 in Dong et al., 2003) as its closest homologues. In grapevine, numerous partial sequences from EST databases showing significant homologies with WRKY genes have been identified from libraries constructed after abiotic stresses or from different stages of flower and fruit development. However, no functional analyses of genes encoding these sequences have been published to date.
Consistent with its putative role as a transcription factor, the VvWRKY1 protein contains a nuclear targeting sequence. Moreover, since phosphorylation of WRKY proteins is thought to play a role in their activation (Yang et al., 1999; Yamamoto et al., 2004), the presence of phosphorylation sites in the VvWRKY1 deduced protein sequence was sought. Indeed, numerous potential phosphorylation sites were found using the PSORT program (Nakai and Kanehisa, 1992; http://psort.nibb.ac.jp) (data not shown). This suggests a possible regulation of VvWRKY1 activity by phosphorylation via different protein kinases.
WRKY proteins were described as transcription factors capable of binding W-box-containing sequences (Eulgem et al., 2000) that are present in the promoter regions of a large number of defence genes including PR genes. The present data show that the VvWRKY1 protein also binds specifically to W-box cis-elements in different nucleotidic environments. Thus, one must assume that VvWRKY1 might regulate the expression of genes containing the W-box within their promoters.
WRKY genes have been duplicated many times over during plant evolution, resulting in a large gene family involved in regulating a large set of genes and thereby ensuring proper cellular responses to physiological processes, internal and external stimuli. For example, the expression of several Arabidopsis WRKY genes is strongly up-regulated during plant senescence (Hindehofer and Zentgraf, 2001; Robatzek and Somssich, 2001; Chen et al., 2002; Guo et al., 2004). In the present study, VvWRKY1 appeared to be developmentally regulated in leaves and in berries. VvWRKY1 expression was very weak in berries before veraison but there was a significant increase at veraison that continued through ripening (Fig. 4A). This finding is in agreement with previous results obtained with the CaWRKY1 gene in pepper. CaWRKY1 expression is strongly up-regulated in red fruit and may play an important role in pepper fruit maturation (Ulker and Somssich, 2004). Furthermore, VvWRKY1 expression in grape berries appears to be co-regulated with some PR-like class genes, such as class IV chitinase, thaumatin-like protein, lipid transfer protein, and metallothionein, that are differentially expressed during ripening, some peaking after veraison (Robinson et al., 1997; Tattersall et al., 1997; Salzman et al., 1998; Davies and Robinson, 2000). Even if some of these genes are induced by pathogen infection in a susceptible cultivar (Jacobs et al., 1999), up-regulation of PR genes seems to be correlated with resistance to powdery mildew (Chellemi and Marois, 1992).
The response to pathogens is regulated by multiple signal transduction pathways in which SA, jasmonic acid, and ethylene function as key signalling molecules (Glazebrook, 2001). SA is a key endogenous secondary signal involved in activation and/or potentiation of plant defence responses (Dempsey et al., 1999). It is required for the establishment of SAR. A second signalling pathway, which is generally antagonistic with the SA-dependent pathway, involves methyl jasmonate/ethylene as key intermediates. Ethylene plays a role in controlling symptom development due to virulent bacteria or fungi (Lund et al., 1998). However, simultaneous activation of these two pathways is fully compatible in Arabidopsis (Van Wees et al., 2000). H2O2 is also an important component of plant defence mechanisms even if its signalling role is not completely defined (Tenhaken et al., 1995). All these key molecules act in a very complex signalling network and share several steps or enzymes with each other (Reymond and Farmer, 1998). In the present assays, all of these molecules affected VvWRKY1 expression, with transcription of VvWRKY1 being up-regulated in response to SA, H2O2, and ethephon. The closest homologues of VvWRKY1, StWRKY1 and AtWRKY75, have also been shown to be induced by elicitors, SA treatment, and pathogen infection (Dellagi et al., 2000; Dong et al., 2003). Moreover, numerous studies showed induction of WRKY gene expression in response to SA (Yang et al., 1999; Chen and Chen, 2000, 2002; Dong et al., 2003; Li et al., 2004), but few authors analysed the effect of other signal molecules, such as H2O2 (Miao et al., 2004; Rhizhsky et al., 2004) or ethylene (Li et al., 2004). Only AtWRKY70 expression can be activated by SA and ACC (a natural precursor of ethylene; Li et al., 2004). On the other hand, no variation in expression of GaWRKY1 was noted by Xu et al. (2004) in response to SA and H2O2. VvWRKY1 activation of transcription was also previously described in grapevine cell suspension after treatment by ergosterol, a non-specific fungal elicitor (Laquitaine et al., 2006). Taken together, these results are consistent with the finding that expression of many WRKY genes in various plant species is rapidly activated upon pathogen challenge and/or elicitor treatments. The induction of VvWRKY1 transcription by wounding, fungal elicitors, or signalling molecules strongly suggests the involvement of this transcription factor in grapevine stress responses.
In this work, an attempt was also made to analyse the biological impact of the constitutive expression of VvWRKY1 in tobacco. All independent transgenic tobacco lines showed no visible phenotypic changes, which is consistent with the present finding that these lines do not have constitutive high elevated levels of PR gene transcripts. This last finding was quite surprising because overexpression of WRKY genes often induced constitutive PR gene expression (Chen and Chen, 2002; Li et al., 2004; Liu et al., 2006). However, the present analysis of PR gene expression was not exhaustive and the possibility cannot be ruled out that the expression of other PR genes can be induced in the transgenic lines. Nevertheless, evidence presented here shows that overexpression of VvWRKY1 in tobacco results in a slight but significant decrease in susceptibility towards three different fungi, namely Pythium, and the downy and the powdery mildew agents. These findings indicate that constitutive ectopic expression of VvWRKY1 alone is insufficient to activate defence responses but requires additional unknown pathogen-induced plant components to exert its functions. Similar results have been obtained in Arabidopsis for AtWRKY18, which alone is not able to activate PR gene expression in transgenic plants during the early stages of development (Chen and Chen, 2002). Like many other complex biological processes, plant defence responses to pathogen infection involve transcriptional regulation of a large number of plant host genes (Rushton and Somssich, 1998). Taken together, these results suggest that the VvWRKY1 transcription factor requires co-ordination with developmentally regulated components and/or other transcription factors induced by plant–pathogen interaction in order to activate plant defence responses in tobacco.
In conclusion, a new grape WRKY gene, designated VvWRKY1, has been identified whose expression is strongly regulated during berry and leaf development, and under stress conditions. This transcription factor appears to be induced by abiotic and biotic stresses in grapevine. Its overexpression in tobacco induces a decrease in susceptibility towards some fungi, suggesting a role in the plant defence response to fungal pathogens. However, the decrease in susceptibility is more significant against a necrotrophic fungus (Pythium) and a biotrophic fungus (powdery mildew) than against the two other biotrophic pathogens tested (downy mildew and PVY). Mechanisms of plant defence against biotrophic and necrotrophic pathogens use different signalling pathways that activate various responses allowing resistance (Glazebrook, 2005). In the case of biotrophs, the HR, often mediated by SA signalling, generally results in resistance, while resistance to necrotrophs involves a different set of defence responses activated by jasmonic acid and/or ethylene signalling. The data presented here suggest that VvWRKY1 may be involved in both signalling pathways. Consistent with this finding, AtWRKY70 has also been shown to be involved in defence against biotrophic and necrotrophic pathogens (AbuQamar et al., 2006; Li et al., 2006). However, the difference in susceptibility observed between VvWRKY1 tobacco transgenic lines and control plants towards all the selected fungi is relatively slight and may be due to the use of a heterologous plant system. Further investigations using a homologous expression system in which VvWRKY1 will find its endogenous target genes and/or partners are in progress and might provide new insights into the biological function of WRKY genes in grapevine and their role in enhancing protection against various fungi.
We acknowledge Dr Y Marco and Dr I Somssich for comments and advice on the manuscript. This work was supported by grants from the ‘Conseil Interprofessionnel du Vin de Bordeaux’ (CIVB).
References
Author notes
These authors contributed equally to this work.
Deceased during the evaluation of this manuscript.





Comments